Genetics, Genomics, and Epigenetics
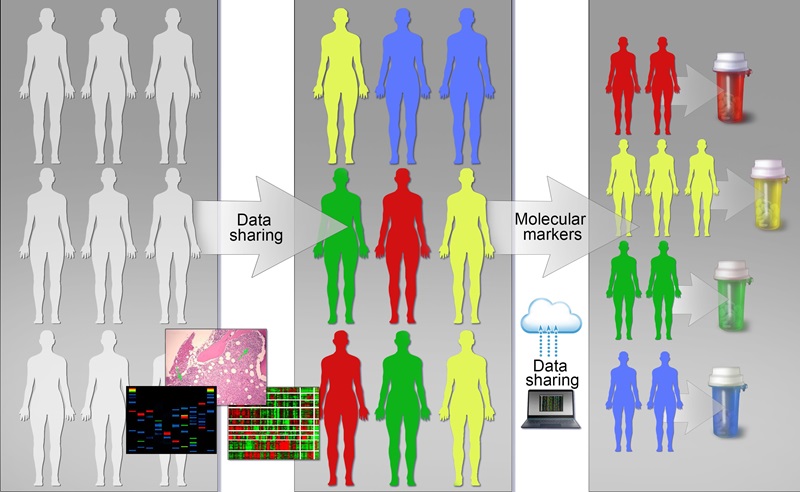
Gene discovery science has been a rich source of innovation in hematology, and hematology has been an ideal testing ground for technologic innovation in this area. Hematology has advanced genetics, genomics, and epigenetics through increased application of technologies in precision medicine, including mutational analyses, epigenetic modulation and gene control, and transcriptomics. Continued development and use of new analytic platforms including single cell sequencing, epigenomics, and proteomics will further broaden this area and its impact on hematology and patient outcomes.
- Increase use and integration of genomic and epigenetic data with clinical parameters to identify new biomarkers of disease and to apply this to the clinical context
- Develop risk models that better reflect our understanding of the genomic basis of hematologic diseases and use of functional genomic tools in these models for screening and treatment stratification
- Investigate the interrelationship between genetic alterations in the tumor cell and the immune microenvironment to identify therapeutic implications that link genetics/epigenetics to immunology/immunotherapy