Genome Editing and Cellular Therapy
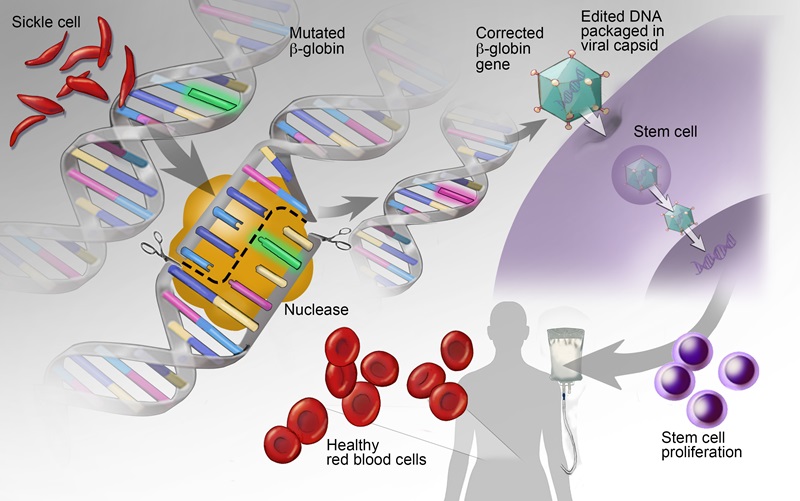
Advances in gene editing and cellular therapies offer promising new therapeutic options that are poised to revolutionize care for those with hematologic conditions. We are at the forefront of understanding how to integrate genomic data with technologies to alter gene expression and cellular function to combat and eliminate disease.
- Enhance use of gene editing for basic/discovery science to provide a pathway that translates genomic data into actionable therapies
- Develop and extend genomic and proteomic approaches to identify and credential novel therapeutic targets for gene therapy, cellular therapy, and cell surface-targeting therapeutic modalities
- Develop novel endpoints for clinical trials that employ gene-editing and cellular therapy
- Expand approaches and application of cellular therapies, including new technologies, targets, modalities, and clinical contexts where cellular therapy can achieve clinical efficacy in malignant and non-malignant disease